|
Building a simple polymer (using the OPLS force field)
(click below for input files)
|
|
 |

|
 |

|
 1ns 1ns |
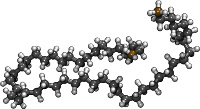
|
|
ch3group.lt
|
|
ch2group.lt
|
|
alkane50.lt
system.lt
(Note: more complex shapes are possible)
|
|
README.TXT
run.in.min
run.in.nvt
|
| Build Using: |
moltemplate.sh system.lt
(warnings and recommendtions...)
|
| Run Using: |
lmp_mpi -i run.in.min
lmp_mpi -i run.in.nvt
|
|
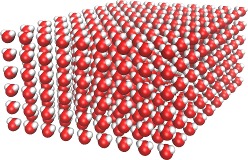
Carbon-Nanotube capillary
(all-atom, explicit water)
(click below for input files)
|
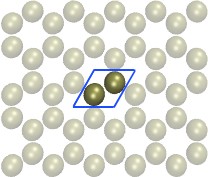
|

|
 (reader's comment:
atoms near junction should be adjusted.
Chiral nanotubes need special treatment.)
(reader's comment:
atoms near junction should be adjusted.
Chiral nanotubes need special treatment.)
|

|

|
 |
 ....
....
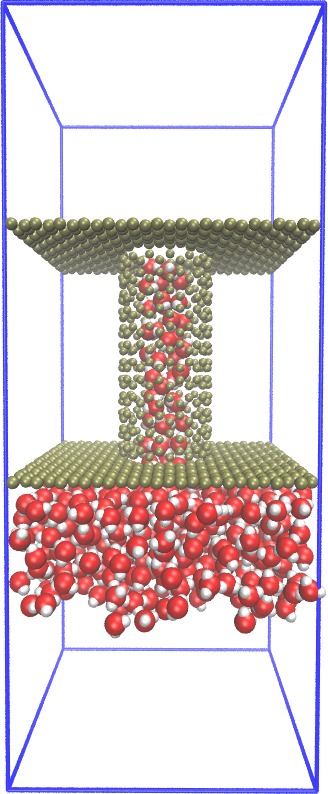
|
graphene.lt
|
|
nanotube.lt,
graphene_walls.lt
|
|
spce.lt,
water_box.lt
|
|
system.lt,
run.in.nvt,
(video)
|
| Build Using: |
moltemplate.sh system.lt
|
| Run Using: |
lmp_mpi -i run.in.nvt
|
|
Aggregation of Simple Toy Polymers (coarse grained)
(click below for input files) |

|
 |
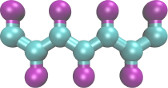
(Note: more complex shapes are possible)
|
 |
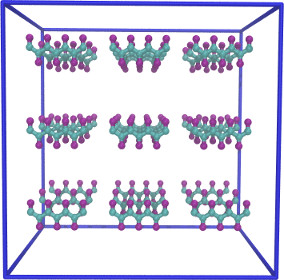
|
 5x10^6 steps 5x10^6 steps |

|
monomer.lt
forcefield.lt
|
|
polymer.lt
|
|
system.lt
|
|
run.in.nvt
|
| Build Using: |
moltemplate.sh -atomstyle "full" system.lt
|
| Run Using: |
lmp_mpi -i run.in.nvt
|
|
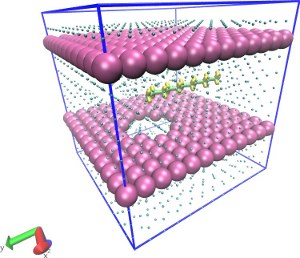
Translocation of a short polymer through a pore
(Explicit LJ Solvent)
(click below for input files)
|
| Requirements: |
This example requires that LAMMPS is
built with
the optional
RIGID
package. |
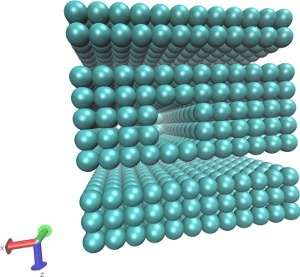
|

|

|
 |
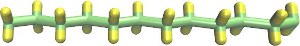
|
 |
|
solvent_single.lt,
solvent.lt
|
|
wall_single.lt,
walls.lt
|
|
monomer.lt,
polymer.lt,
polymer_forcefield.lt
|
|
system.lt,
run.in.npt,
(video)
|
| Build Using: |
moltemplate.sh system.lt
|
| Run Using: |
lmp_mpi -i run.in.npt
|
|
Coarse-grained lipid bilayer
using the MARTINI force field and PACKMOL
(click below for input files)
|
| Requirements: |
This example uses PACKMOL
(For an alternative method, see system.lt )
|



|  |

|
 13ns 13ns |

|
 13ns 13ns |

|
README.TXT
lipid.lt
water.lt
system.lt
|
|
README_pm.txt
lipid.xyz
water.xyz
mix_lipids+water.inp
|
|
run.in.min
run.in.npt
run.in.nvt
|
|
(video)
|
| Build Using: |
packmol < mix_lipids+water.inp
moltemplate.sh -xyz system.xyz system.lt
|
| Run Using: |
lmp_mpi -i run.in.min
lmp_mpi -i run.in.npt
lmp_mpi -i run.in.nvt
|
|
Many-Body force field example: mW solvent + CG hydrocarbon mixture
(Many-body force fields can be combined with ordinary, pairwise-additive force fields. Click below for input files.)
|
| Requirements: |
This example requires that LAMMPS is
built with
the
MANYBODY
package. |

|

|

|
 |

|
 400 ps 400 ps |

|
README.png
watmw.lt
|
|
cyclododecane.lt,
trappe1998.lt
|
|
README.TXT
system.lt
|
|
run.in.npt
(video)
|
| Build Using: |
moltemplate.sh -a "@atom:/WatMW/mW 1" system.lt
|
| Run Using: |
lmp_mpi -i run.in.npt
|
|
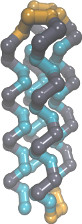
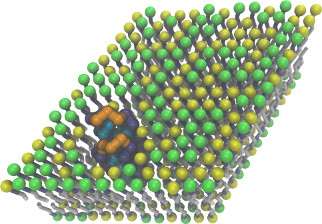
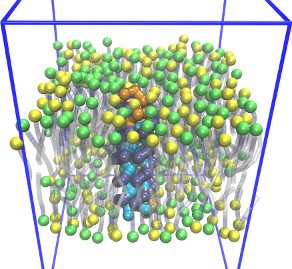
Coarse-grained membrane protein
click below for input files
|
| Requirements: |
This example requires that LAMMPS is
built with
the optional
USER-MISC
package, before
additional code
is added (in that order)
|



|

|
|
 |
|
 3ns 3ns |
|
CGLipidBr2005.lt,
table_int.dat
|
|
1beadProtSci2010.lt,
|
|
system.lt
|
|
run.in.npt
|
| Build Using: |
moltemplate.sh system.lt
|
| Run Using: |
lmp_mpi -i run.in.npt
|
|
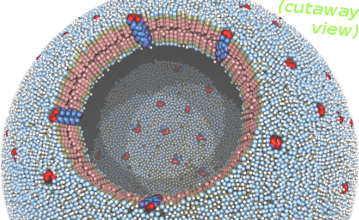
Vesicle with protein inclusions
click below for input files
|
| Requirements: |
This example requires PACKMOL.
LAMMPS must be
built with
the optional
USER-MISC
package, before
additional code
is added (in that order)
|



|
 |
|
Moltemplate files:
README_mt.txt,
CGLipidBr2005.lt,
table_int.dat,
1beadProtSci2010.lt,
system.lt
|
|
PACKMOL files:
README_pm.txt,
DPPC.xyz,
protein.xyz,
step1_proteins.inp,
step2_innerlayer.inp,
step3_outerlayer.inp
|
|
LAMMPS files:
run.in.min,
run.in.make_uniform
run_T=345K.in
This is a complex example requiring hours or days to set up.
Please follow the instructions in the README files.
|
| Build Using: |
packmol < step1_proteins.inp # requires ~40 minutes
packmol < step2_innerlayer.inp # requires ~10 hours
packmol < step3_outerlayer.inp # requires 1-3 days (creates system.xyz)
moltemplate.sh -xyz system.xyz system.lt
|
| Run Using: |
lmp_mpi -i run.in.min
lmp_mpi -i run.in.make_uniform
lmp_mpi -i run_T=345K.in
|
|
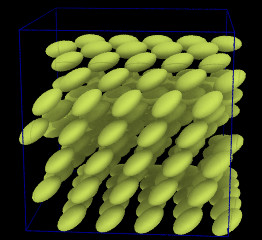
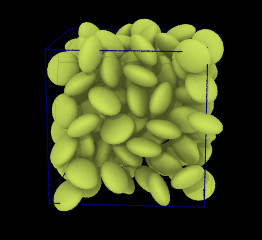
Ellipsoidal particles
(Moltemplate can build systems containing ellipsoids and point-dipoles.
Click below for input files)
|
| Requirements: |
This example requires that LAMMPS is
built with
the optional
ASPHERE
package. |

|
 |
|
 |
|
|
|
README.txt
benzene_cg.lt
|
|
system.lt
|
|
run.in
|
|
|
| Build Using: |
moltemplate.sh -atomstyle "atomid atomtype flag density x y z" system.lt
-allow-wildcards -nocheck
|
| Run Using: |
lmp_mpi -i run.in
|
 )
)